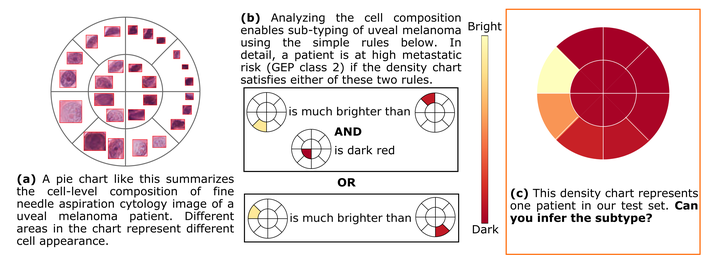
 An overview of the algorithm.
An overview of the algorithm.
Abstract
Algorithmic decision support is rapidly becoming a staple of personalized medicine, especially for high-stakes recommendations in which access to certain information can drastically alter the course of treatment, and thus, patient outcome; a prominent example is radiomics for cancer subtyping. Because in these scenarios the stakes are high, it is desirable for decision systems to not only provide recommendations but supply transparent reasoning in support thereof. For learning-based systems, this can be achieved through an interpretable design of the inference pipeline. Herein we describe an automated yet interpretable system for uveal melanoma subtyping with digital cytology images from fine needle aspiration biopsies. Our method embeds every automatically segmented cell of a candidate cytology image as a point in a 2D manifold defined by many representative slides, which enables reasoning about the cell-level composition of the tissue sample, paving the way for interpretable subtyping of the biopsy. Finally, a rule-based slide-level classification algorithm is trained on the partitions of the circularly distorted 2D manifold. This process results in a simple rule set that is evaluated automatically but highly transparent for human verification. On our in house cytology dataset of 88 uveal melanoma patients, the proposed method achieves an accuracy of 87.5% that compares favorably to all competing approaches, including deep “black box” models. The method comes with a user interface to facilitate interaction with cell-level content, which may offer additional insights for pathological assessment.